Histone Proteins | Vibepedia
Histone proteins are fundamental to eukaryotic life, acting as the spools around which DNA winds to form nucleosomes, the basic units of chromatin. These…
Contents
- 🎵 Origins & History
- ⚙️ How It Works
- 📊 Key Facts & Numbers
- 👥 Key People & Organizations
- 🌍 Cultural Impact & Influence
- ⚡ Current State & Latest Developments
- 🤔 Controversies & Debates
- 🔮 Future Outlook & Predictions
- 💡 Practical Applications
- 📚 Related Topics & Deeper Reading
- Frequently Asked Questions
- References
- Related Topics
Overview
The story of histones begins not with a single eureka moment, but with decades of meticulous biochemical investigation into the composition of chromosomes. Early work in the late 19th and early 20th centuries identified proteins as major components of the nucleus, but their specific roles remained elusive. By the 1940s, researchers like Ivan Tolmach were beginning to characterize the physical properties of these nuclear proteins, noting their basic nature. The concept of DNA as genetic material gained traction in the 1950s, following the work of James Watson and Francis Crick, which spurred intense interest in how DNA was organized. The pivotal discovery of the nucleosome structure, the fundamental unit of chromatin, is largely credited to Roger D. Kornberg, whose seminal work in the early 1970s, published in journals like Nature, revealed that DNA was wrapped around a core of histone proteins. This breakthrough, building on earlier observations by Albert L. Dounce, fundamentally reshaped our understanding of genome organization and regulation, earning Kornberg the Nobel Prize in Physiology or Medicine in 2006.
⚙️ How It Works
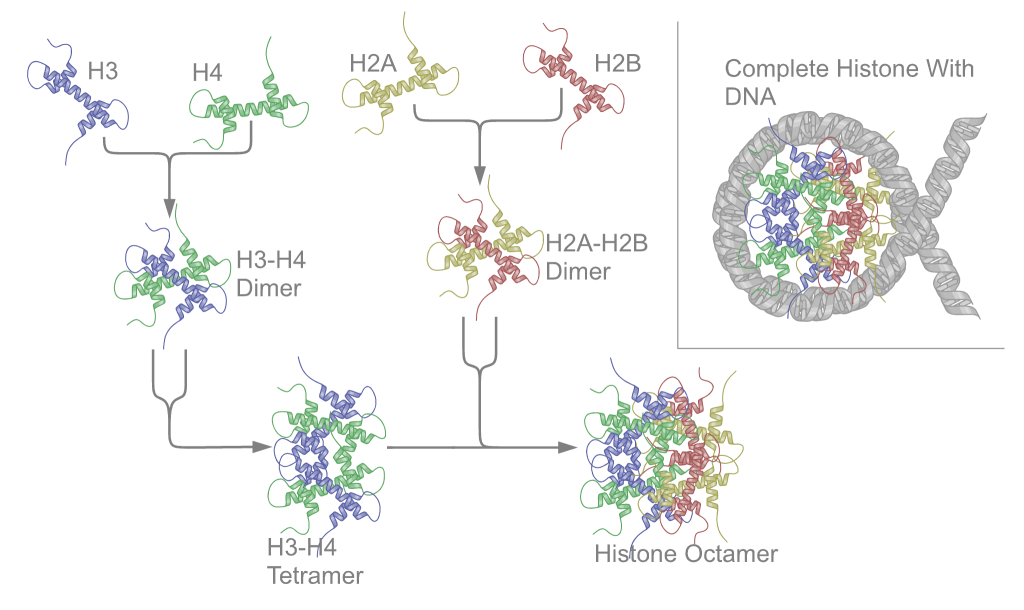
Histone proteins function as molecular spools, enabling the astonishing feat of packaging approximately 2 meters of human DNA into a nucleus only about 10 micrometers in diameter. The core histones—two copies each of H2A, H2B, H3, and H4—assemble into an octameric core particle. DNA then wraps around this histone octamer approximately 1.65 times, forming a nucleosome, which is the fundamental repeating unit of chromatin. This structure is further stabilized by linker histones, primarily H1, which bind to the DNA where it enters and exits the nucleosome, facilitating the formation of higher-order chromatin structures like the 30-nanometer fiber. This compacting process is not static; it's a dynamic interplay where chemical modifications to histone tails—such as acetylation, methylation, and phosphorylation—act as signals that can either promote or repress gene expression by altering chromatin accessibility. These modifications are orchestrated by a complex network of enzymes, including histone acetyltransferases (HATs) and histone deacetylases (HDACs), and are crucial for processes like transcription, DNA repair, and cell cycle progression.
📊 Key Facts & Numbers
A typical human cell contains roughly 30 million nucleosomes, each comprising about 147 base pairs of DNA wrapped around a histone octamer. The total mass of DNA in a human cell is approximately 6 picograms, and histones constitute about 50% of the mass of chromatin. There are five main families of histones: H1, H2A, H2B, H3, and H4, with H2A, H2B, H3, and H4 forming the core nucleosome. The human genome encodes at least 70 different histone variants, each potentially conferring unique structural or regulatory properties. For instance, the histone variant H3.3 is incorporated during DNA replication and transcription to maintain active gene expression. The precise stoichiometry of these proteins is critical; a deviation of even a few percent can lead to cellular dysfunction and disease. The sheer scale of this packaging is staggering: if stretched out, the DNA from a single human cell would measure about 1.8 meters, but when packaged by histones, it fits within a nucleus less than 0.01 millimeters in diameter.
👥 Key People & Organizations
The foundational work on nucleosome structure was pioneered by Roger D. Kornberg, whose research in the 1970s elucidated the basic unit of chromatin. Albert L. Dounce was an early investigator of nuclear proteins, proposing their role in chromosome structure in the 1940s. More recently, Allan R. Campbell and Michael Grunstein have made significant contributions to understanding histone modifications and their role in gene regulation, with Grunstein’s lab at UCLA being a powerhouse in epigenetics research. Organizations like the American Society for Cell Biology and the Genetics Society of America foster research and disseminate findings on histones. Major research institutions such as The Rockefeller University and MIT host leading labs investigating histone biology and its implications for human health.
🌍 Cultural Impact & Influence
Histones have profoundly influenced our understanding of genetics and cell biology, moving the field beyond a purely DNA-centric view to embrace the crucial role of epigenetics. The discovery of nucleosomes and histone modifications has permeated textbooks and undergraduate curricula, becoming a cornerstone of molecular biology education. This understanding has fueled the development of new research tools and techniques, such as chromatin immunoprecipitation (ChIP), which allows scientists to map histone modifications and protein binding sites across the genome. The concept of histones as dynamic regulators has also captured the public imagination, appearing in popular science books and documentaries that explore the complexities of heredity and gene expression. Furthermore, the medical implications of histone dysregulation have spurred public interest in epigenetics as a potential avenue for treating diseases like cancer and neurological disorders.
⚡ Current State & Latest Developments
The current frontier in histone research involves unraveling the precise mechanisms by which histone variants and their modifications contribute to cellular identity and disease. Researchers are actively developing sophisticated techniques, including cryo-electron microscopy and single-cell multi-omics, to visualize chromatin structure and epigenetic states at unprecedented resolution. The development of targeted therapies that modulate histone-modifying enzymes, such as Vorasidenib (a dual HDAC inhibitor approved for certain brain tumors), represents a major leap forward. The field is also exploring the role of histones in non-coding DNA regions and their involvement in complex traits and developmental processes. The integration of artificial intelligence and machine learning is accelerating the analysis of vast epigenomic datasets, promising to uncover novel regulatory networks and therapeutic targets.
🤔 Controversies & Debates
A significant debate revolves around the precise functional impact of specific histone variants and the combinatorial effects of multiple histone modifications. While the general principles of histone acetylation promoting gene expression and methylation having diverse roles are established, the specificity and context-dependency of these marks remain areas of active investigation. Some researchers argue that the 'histone code'—the idea that specific patterns of modifications dictate distinct cellular outcomes—is overly simplistic, emphasizing the complex interplay with other regulatory factors. Another point of contention is the therapeutic window for histone-modifying drugs; while promising, these drugs can have significant off-target effects and toxicities, leading to ongoing efforts to develop more precise interventions. The role of histone chaperones in nucleosome assembly and disassembly also presents complex questions regarding their specificity and regulation.
🔮 Future Outlook & Predictions
The future of histone research points towards increasingly personalized epigenetic therapies. As our understanding of the histone code deepens, we can expect the development of highly specific drugs targeting particular histone modifiers or variants for diseases ranging from cancer to neurodegenerative disorders. The integration of epigenomic data with genomics and transcriptomics will likely lead to more accurate disease prognostics and diagnostics. Furthermore, advances in gene editing technologies, such as CRISPR-Cas9, may eventually allow for direct manipulation of histone genes or their regulatory elements to correct epigenetic defects. The potential to reprogram cell fate by altering histone landscapes could revolutionize regenerative medicine, enabling the generation of specific cell types for therapeutic purposes. We may also see the development of epigenetic clocks based on histone modifications for more accurate aging assessments.
💡 Practical Applications
Histone proteins are central to several critical practical applications in biotechnology and medicine. Epigenetic drugs, particularly histone deacetylase (HDAC) inhibitors like Vorasidenib and Romidepsin, are already used in cancer therapy to reactivate silenced tumor suppressor genes. Histone modifications are also being explored as biomarkers for disease diagnosis and prognosis, particularly in oncology. In research, techniques like ChIP-sequencing allow scientists to map regulatory elements and understand gene expression patterns, aiding in drug discovery and development. Furthermore, understanding histone function is crucial for developing cell-based therapies, such as reprogramming somatic cells into induced pluripotent stem cells (iPSCs), a process heavily reliant on epigenetic remodeling. The development of diagnostic tools that can detect aberrant histone patterns could also lead to earlier disease detection.
Key Facts
- Year
- 1970s (nucleosome discovery)
- Origin
- Eukaryotic cells
- Category
- science
- Type
- concept
Frequently Asked Questions
What exactly are histone proteins and what do they do?
Histone proteins are highly basic proteins found in eukaryotic cell nuclei. Their primary role is to act as spools around which DNA winds to form nucleosomes, the fundamental units of chromatin. This packaging compacts the vast length of DNA, making it fit within the nucleus and protecting it from damage. Beyond structural roles, histones are crucial for regulating gene expression through chemical modifications.
How do histones influence gene expression?
Histones influence gene expression through a process called epigenetics. Chemical modifications to histone tails, such as acetylation and methylation, alter the structure of chromatin. For example, histone acetylation generally loosens chromatin, making genes more accessible to transcription machinery and thus promoting gene expression. Conversely, certain methylation patterns can lead to gene silencing. This dynamic modification system allows cells to control which genes are turned on or off.
What are the different types of histones?
There are five main families of histones: H1, H2A, H2B, H3, and H4. H2A, H2B, H3, and H4 are known as core histones and form the octameric core around which DNA wraps to create a nucleosome. H1, the linker histone, binds to the DNA where it enters and exits the nucleosome, playing a role in higher-order chromatin compaction. Additionally, there are numerous histone variants within these families that can confer specialized functions.
Why are histones important for human health?
Histones are vital for maintaining genome stability and regulating gene expression, processes fundamental to all cellular functions. Dysregulation of histone proteins and their modifications is implicated in numerous diseases, most notably cancer, where it can lead to uncontrolled cell growth and proliferation. Aberrant histone activity is also linked to neurological disorders, developmental abnormalities, and aging. Consequently, histones and the enzymes that modify them are significant targets for therapeutic interventions.
Can histone function be therapeutically targeted?
Yes, histone function is a major target for drug development, particularly in oncology. Drugs known as epigenetic modifiers, such as histone deacetylase (HDAC) inhibitors (e.g., Vorasidenib) and histone methyltransferase inhibitors, are used to treat certain cancers. These drugs aim to restore normal gene expression patterns by altering histone modifications, thereby inhibiting tumor growth or reactivating silenced tumor suppressor genes. Research is ongoing to develop more specific and less toxic histone-targeting therapies.
How much DNA can histones pack into a cell?
Histone proteins are incredibly efficient at packaging DNA. In a human cell, the approximately 1.8 meters of DNA, if stretched out, can be compacted by histones into a nucleus measuring only about 10 micrometers in diameter. This remarkable feat is achieved through the formation of nucleosomes and subsequent higher-order chromatin structures, reducing the DNA length by a factor of roughly 180,000.
What is the 'histone code' hypothesis?
The 'histone code' hypothesis proposes that specific patterns of histone modifications act as a language, dictating distinct downstream biological outcomes, such as gene activation or repression. It suggests that combinations of different marks on histone tails are read by specific proteins, leading to defined cellular responses. While the concept has been influential, the precise interpretation and combinatorial complexity of this 'code' remain subjects of ongoing research and debate, with many arguing for a more context-dependent and less deterministic interpretation.